Two primers for the target site 1 and 2 primers for the target size 2 (see image below): To generate 2 sgRNAs with each their target site (gRNA1 and gRNA2 indicated by blue arrows and highlighted in yellow in image below), 4 primers need to be designed. They can target 2 different regions of the genome or target the same gene for instance. The vector pCBC-DT1T2 contain 2 sgRNAs, therefore 2 target sequences. This file can be open with the free software SnapGene viewer (download here). Plasmid available in genebank format in plasmids/pHEE401E.gbk. It contains the complementary parts of the pCBC-DT1T2 which ensures the proper expression of the 2 sgRNAs plus other features needed for Agrobacterium transformation and selection. 
PHEE401E is based on pCambia backbone for Agrobacterium-mediated transformation. 
These sequences are actually in the pHEE401E binary vector and will be integrated in place after the digestion with BsaI enzyme and the ligation (GoldenGate cloning). Note that the vector pCBC-DT1T2 does not contain the promoter of the first sgRNA and the second sgRNA contains neither its scaffold sequence nor its terminator. BsaI restriction sites are highlighted in yellow. Part of the plasmid pCBC-DT1T2 containing the elements which will be transfered into pHEE401E during the GoldenGate cloning. T4 ligase is used to ligate the corresponding cohesive ends (it is also able to ligate blunt ends but with much lower efficiency). The 5 nucleotides overhang can be chosen and included in any construct to allow ligation. The enzyme cuts 1 nucleotide directly upstream of the recognition site on the + strand but 5 nucleotides upstream on the - strand (highlighted in yellow in figure below). The main principle to understand GoldenGate cloning is that a type IIS restriction enzyme such as BsaI, used here, is able to cut besides its actual recognition site, allowing to create non-palindromic overhangs which allow directional cloning. See more details about Golden Gate assembly on NEB website. More target sequences can be added by adding primers with specific overhanging sequence for Golden Gate assembly. Plasmid available in genebank format in plasmids/pCBC_DT1T2.gbk. The terminator of the Pisum sativum rbcS E9 gene was tested according to previous observation ( Sarrion-Perdigones et al 2013) and found more efficient than coupled with a NOS terminator. This tissue-specific expression of Cas9 allows to obtain T1 homozygous or biallelic plants instead of mosaic plants. Secondly, a restriction-ligation step is performed to incorporate a part of pCBC-DT1T2 into pHEE401E, removing in the meantime its spectinomycin (SpR) resistance cassette of the latter Xing et al 2014.Ĭas9 expression is driven by the promoter of the egg cell-specific EC1.1 ( AT1G76750) gene and the enhancer EC1.2 gene ( AT2G21740). 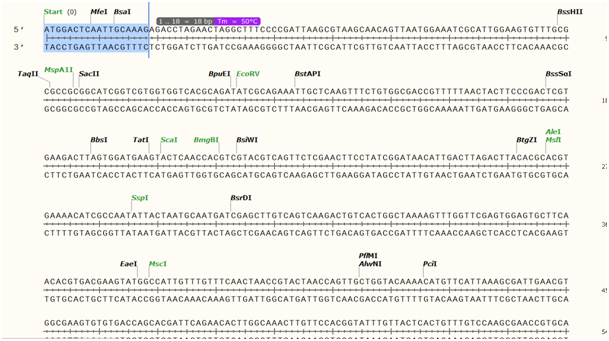
First a PCR on pCBC-DT1T2 allows to add the proper target sequences on the gRNA and the BsaI restriction sites on pCBC-DT1T2.The construct is split into one module vector set (pCBC-DT1T2) containing two single guide RNAs (sgRNAs) and a binary vector based on pCAMBIA (pHEE401E) containing additional sequences ensuring proper expression of the 2 sgRNAs and transformation into Arabidopsis. Since the term target sequence/site sounds more explicit than spacer to define the 19-20 nt sequence which matches the region of the genome to be targeted, we will stick to the target sequence/site across the text. Protospacer adjacent motif (PAM) defines the 3-nt motif adjacent (in 5') to the original protospacer. It becomes the spacer once in the repeat-spacer array and becomes the crRNA when transcribed. Note: the original target sequence in the invading phage which is incorporated into the repeat-spacer array (also called CRISPR array) in bacteria is called the protospacer. Hairpin structures within the sgRNA mimics the crRNA-tracrRNA complex enabling the binding of Cas9. They were merged into one molecule called sgRNA to facilitate cloning. In nature, these 2 elements form 2 separate RNA molecules which hybridize to ensure Cas9 activity at the target site specified by the crRNA. a trans-activating crRNA (tracrRNA), also called scaffold sequence, which binds to the Cas9 nuclease.a CRISPR RNA (crRNA) which contains the target site (also called spacer) of the genome and is usually 15-20 bp long.Table of contents generated with markdown-toc Introduction CRISPR/Cas system single guide RNAĪ single guide RNA (sgRNA) is an engineered RNA molecule containing 2 elements: Cloning and transformation in Arabidopsis.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
